tick.hawkes.HawkesADM4¶
- class tick.hawkes.HawkesADM4(decay, C=1000.0, lasso_nuclear_ratio=0.5, max_iter=50, tol=1e-05, n_threads=1, verbose=False, print_every=10, record_every=10, rho=0.1, approx=0, em_max_iter=30, em_tol=None)[source]¶
A class that implements parametric inference for Hawkes processes with an exponential parametrisation of the kernels and a mix of Lasso and nuclear regularization
Hawkes processes are point processes defined by the intensity:
\[\forall i \in [1 \dots D], \quad \lambda_i(t) = \mu_i + \sum_{j=1}^D \sum_{t_k^j < t} \phi_{ij}(t - t_k^j)\]where
\(D\) is the number of nodes
\(\mu_i\) are the baseline intensities
\(\phi_{ij}\) are the kernels
\(t_k^j\) are the timestamps of all events of node \(j\)
and with an exponential parametrisation of the kernels
\[\phi_{ij}(t) = \alpha^{ij} \beta \exp (- \beta t) 1_{t > 0}\]In our implementation we denote:
Integer \(D\) by the attribute
n_nodesVector \(\mu \in \mathbb{R}^{D}\) by the attribute
baselineMatrix \(A = (\alpha^{ij})_{ij} \in \mathbb{R}^{D \times D}\) by the attribute
adjacencyNumber \(\beta \in \mathbb{R}\) by the parameter
decay. This parameter is given to the model
- Parameters:
decay :
floatThe decay used in the exponential kernel
C :
float, default=1e3Level of penalization
lasso_nuclear_ratio :
float, default=0.5Ratio of Lasso-Nuclear regularization mixing parameter with 0 <= ratio <= 1.
For ratio = 0 this is nuclear regularization
For ratio = 1 this is lasso (L1) regularization
For 0 < ratio < 1, the regularization is a linear combination of Lasso and nuclear.
max_iter :
int, default=50Maximum number of iterations of the solving algorithm
tol :
float, default=1e-5The tolerance of the solving algorithm (iterations stop when the stopping criterion is below it). If not reached it does
max_iteriterationsverbose :
bool, default=FalseIf
True, we verbose thingsn_threads :
int, default=1Number of threads used for parallel computation.
if
int <= 0: the number of physical cores available on the CPUotherwise the desired number of threads
print_every :
int, default=10Print history information when
n_iter(iteration number) is a multiple ofprint_everyrecord_every :
int, default=10Record history information when
n_iter(iteration number) is a multiple ofrecord_every- Attributes:
n_nodes :
intNumber of nodes / components in the Hawkes model
baseline :
np.array, shape=(n_nodes,)Inferred baseline of each component’s intensity
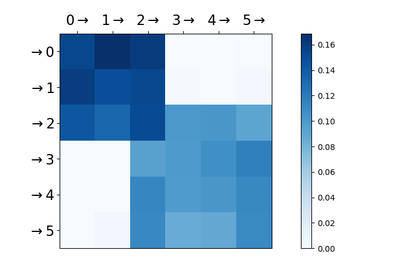
adjacency :
np.ndarray, shape=(n_nodes, n_nodes)Inferred adjacency matrix
References
Zhou, K., Zha, H., & Song, L. (2013, May). Learning Social Infectivity in Sparse Low-rank Networks Using Multi-dimensional Hawkes Processes. In AISTATS (Vol. 31, pp. 641-649).